Single-cell transcriptomics of the human parasite Schistosoma mansoni first intra-molluscan stage reveals tentative ... - Nature.com
Abstract
Schistosomiasis is a major Neglected Tropical Disease, caused by the infection with blood flukes in the genus Schistosoma. To complete the life cycle, the parasite undergoes asexual and sexual reproduction within an intermediate snail host and a definitive mammalian host, respectively. The intra-molluscan phase provides a critical amplification step that ensures a successful transmission. However, the cellular and molecular mechanisms underlying the development of the intra-molluscan stages remain poorly understood. Here, single cell suspensions from S. mansoni mother sporocysts were produced and sequenced using the droplet-based 10X Genomics Chromium platform. Six cell clusters comprising two tegument, muscle, neuron, parenchyma and stem/germinal cell clusters were identified and validated by in situ hybridisation. Gene Ontology term analysis predicted key biological processes for each of the clusters, including three stem/germinal sub-clusters. Furthermore, putative transcription factors predicted for stem/germinal and tegument clusters may play key roles during parasite development and interaction with the intermediate host.
Similar content being viewed by others
Single-cell atlas of the first intra-mammalian developmental stage of the human parasite Schistosoma mansoni
The stage- and sex-specific transcriptome of the human parasite Schistosoma mansoni

Single-cell transcriptomic analysis of bloodstream Trypanosoma brucei reconstructs cell cycle progression and developmental quorum sensing
Introduction
Schistosomiasis, infection with parasitic flatworms in the genus Schistosoma, remains a major neglected tropical disease with more than 240 million people affected worldwide and more than 700 million at risk of infection in endemic areas1,2. To date, a single drug (Praziquantel) is in use; however, this drug is effective only against adult worms, it does not prevent reinfection and drug resistance may be emerging in the field3. One approach that may lead to novel control strategies is to gain a better understanding of the mechanisms underlying life cycle progression, including the cells and their transcriptomic signatures across developmental stages4,5.
Schistosoma mansoni eggs laid by adult worm pairs dwelling in the portal system of the mammalian host traverse the intestinal wall and pass with the faeces into the environment. In contact with fresh water, from the eggs hatch free-swimming larvae (miracidia) that seek, infect a suitable snail, and transform into mother sporocysts, the first intra-molluscan developmental stage. Within the mother sporocyst, groups of stem cells (historically termed 'germinal cells') start to proliferate and differentiate to develop into daughter sporocysts6. By 5 days post infection (5 dpi), developing daughter sporocysts, initially spherical in shape, start to grow within the brood chamber, elongate and become surrounded by a primitive epithelium derived from the mother sporocyst tegument6. By ~ 15 dpi they have acquired the definitive vermiform shape containing germinal cells closely packed in the medial part of the body6. The mature daughter sporocysts escape from the mother sporocyst, migrate through the snail tissue to the digestive gland area and start to produce whole cercariae from single germinal cells following a second round of embryogenesis7. Therefore, from a single miracidium, hundreds to thousands of clonal human-infective cercariae are produced. Extensive knowledge gathered for several decades through detailed histological, and electron microscopy-based studies paved the way towards an understanding of the parasite progression within the snail8,9,10,11. This knowledge can now be scrutinised using current molecular and 'omics' technologies12. Shining new light on the cellular and molecular basis of this parasite expansion strategy is critical not only to discover novel aspects of trematode developmental biology, but also to reveal targets for control4.
Single-cell transcriptome sequencing (scRNA-seq) has been employed to define cellular subtypes by revealing their specific transcriptional signatures. Compared with so-called 'bulk RNA-seq' studies of whole organisms or tissues, scRNA-seq has exceptional resolving power, being able to detect genes expressed in just a few cells or with low expression levels, but also reveals the stochastic nature of gene expression in individual cells13. Single cell transcriptomics have been used in several systems to understand diverse biological processes, such as cell differentiation, tissue specification and development as well as to generate cell "atlases" based on the scRNA-seq profiles across different tissues14. Studies have employed scRNA-seq in S. mansoni4,5 to generate cell atlases for male and female adult worms15, mixed-sex schistosomula16, the first intra-mammalian developmental stage, and more recently mixed-sex miracidia17. In addition, scRNA-seq in schistosomes has been employed to define and functionally characterise stem cell populations driving the development of both intra-molluscan18 and intra-mammalian stages19.
Important contributions to our current understanding of the developmental biology of schistosome intra-molluscan stages have been made using 'bulk RNA-seq'20. In particular, two germinal cell lineages with distinct proliferation properties had previously been identified and functionally characterised by 'bulk transcriptomics' and RNAi, respectively20. However, there is a critical lack of transcriptomic data and knowledge of gene regulatory networks at the single cell level. Wang and collaborators pioneered work in this area by sequencing 35 sporocyst individual cells, focusing primarily on proliferating stem/germinal cells18 but most other cell types of this lifecycle stage remain uncharacterised. In the present study, we have followed an untargeted approach to characterise the individual transcriptomic signatures of more than 600 cells isolated from cultured mother sporocysts. Each of the tissue types were spatially validated by fluorescence in situ hybridisation (FISH). Furthermore, we have explored aspects of gene expression regulation, including the prediction of promoter motifs as tentative binding sites for transcription factors, in the stem/germinal and tegumental cell populations. This study contributes to the expansion of the currently scarce number single cell datasets for schistosomes5 and reveals key candidate genes involved in the intra-snail developmental phase of this neglected tropical disease pathogen.
Results
Six cell populations identified in the Schistosoma mansoni mother sporocyst
Freshly collected S. mansoni miracidia were transferred into sporocyst media to induce transformation into mother sporocysts. Within the first ~ 16 h most parasites have shed the cilia plates, and their tegument has been remodelled8; however, parasite in-vitro development is not synchronous. Therefore, we decided to culture the mother sporocysts for 5 days (named 'D5 sporocysts') to facilitate the complete transformation of > 95% of the parasites10,21 (Fig. 1A and Supplementary Fig. S1). The D5 sporocysts were collected and processed following the dissociation protocol previously used for schistosomula16, and live cells enriched and quantified using Fluorescence-activated Cell Sorting (FACS). The droplet-based 10X Genomics Chromium platform was used to generate transcriptome-sequencing data from a total of 601 cells after applying quality-control filters. With the assistance of a high-resolution nuclei quantification protocol, based on a machine-learning imaging platform22, we estimated that a D5 sporocyst contains an average of 169 nuclei (n = 5; range: 112–254) (Supplementary Fig. S2). Therefore, the number of quality-controlled cells theoretically represents > 3.5 × coverage of all cells in a single D5 mother sporocyst.
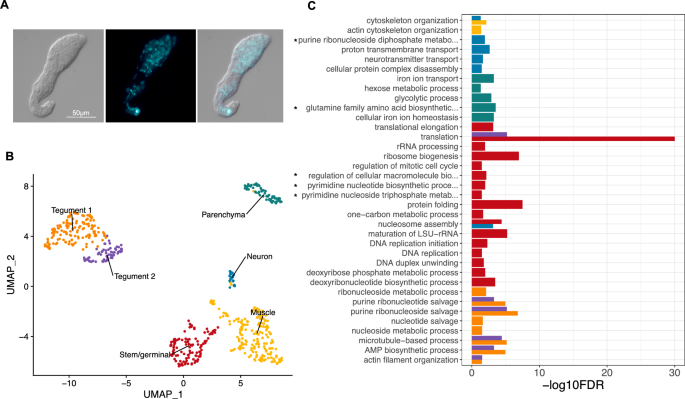
Six distinct cell populations identified in D5 mother sporocysts. (A) Representative picture of D5 sporocyst in DIC bright field (left), DAPI-fluorescent field (centre) and merged (right). Scale bar: 50 μm. (B) Uniform Manifold Approximation and Projection (UMAP) representation of 601 single cells from D5 sporocysts. The cell clusters are coloured and labelled as indicated. The list of the Seurat marker genes in all cell clusters is provided in Supplementary Table S3. (C) Gene ontology (GO) enrichment analysis for biological processes only (marker genes with minimum AUC = 0.7, and GO terms supported by ≥ 2 genes), for top marker genes in each indicated colour-coded cell cluster (as shown in B). Only statistically significant GO enriched biological processes are depicted (− log10 (FDR < 0.05)). FDR: False Discovery Rate. Full names of GO terms indicated with *: purine ribonucleoside diphosphate metabolic process (GO:0009179), glutamine family amino acid biosynthetic process (GO:0009084), regulation of cellular macromolecule biosynthetic process (GO:2000112), pyrimidine nucleotide biosynthetic process (GO:0006221), pyrimidine nucleoside triphosphate metabolic process (GO:0009147). Full data provided in Supplementary Tables S4 and S5, for non-specificity filtering analysis with AUC = 0.7 and AUC = 0.6, respectively.
Based on top markers identified using Seurat, annotation from schistosomula single cell data and genes curated from the literature, we identified six discrete cell populations (Supplementary Table S3); Tegument-1 (138 cells), Tegument-2 (66 cells), Muscle (189 cells), Stem/germinal (119 cells), Parenchyma (66 cells) and Neuron (23 cells) (Fig. 1B). To further explore the biological processes in which the cells of each cluster are involved, we examined over-represented Gene Ontology terms using TopGO (Fig. 1C). Within each cluster, there were clear examples of single genes with annotated roles that are statistically enriched due to their rarity in the genome (Supplementary Tables S4, S5). Although many of these conformed to expectations—for instance, 'neuropeptide signalling' is enriched in the Neuron cluster due to the expression of the known marker gene 7B2—we focussed on over-represented annotation supported by multiple genes. In the case of the Neuron cluster, this highlighted 'neurotransmitter transport', as well as less expected terms 'nucleosome assembly' and 'proton transmembrane transport. In the Muscle cluster, 'cytoskeletal organisation' was enriched, and previously characterised serotonin receptors in schistosomes23 showed muscle-specific expression (Supplementary Fig. S3). The Parenchyma cluster showed enrichment for several aspects of metabolism including, 'iron homeostasis', 'amino acid metabolism' and 'glycolysis'; whereas the Stem/germinal cell cluster, unsurprisingly, was highly enriched for genes involved in DNA replication, ribogenesis and translation.
In both Tegument clusters, 'purine ribonucleotide/side salvage' was highly enriched, as was 'microtubule-based process' due to the expression of 6–8 dyneins and the dynein-domain protein SmTAL2 (Fig. 1C and Supplementary Tables S4, S5). To further explore differences between the two Tegument subclusters, we first identified differentially expressed genes between Tegument 1 and 2 (Supplementary Tables S6, S7), and then performed a GO term enrichment analysis for each of the sub-clusters. 'Proteolysis', 'metalloendopeptidase activity', and 'metal ion binding' were GO terms supported by at least 2 genes found enriched in Tegument 1 (Supplementary Figure S4A and Table S8). On the other hand, Tegument 2 showed a significant enrichment in the GO terms 'translation', 'structural constituent of ribosome', 'ribosome' and 'large ribosomal subunit', i.e., biological processes and cellular components associated with protein synthesis. Remarkably, four out of the top 5 upregulated genes in Tegument 2 compared to Tegument 1 were ribosomal proteins (Supplementary Table S7); however, the proportion of cells in each of the subclusters expressing these genes was similar, e.g., the 40S ribosomal protein S24 is expressed in more than 90% of the cells in both Tegument subclusters (Supplementary Table S7). Most of the genes highly expressed in Tegument 2 were also expressed in Tegument 1, with few exceptions including an arrestin_C domain-containing protein (Smp_121950) expressed in ~ 54 and ~ 74% of the cells in Tegument 1 and 2, respectively (Supplementary Fig. S4B). On the contrary, several genes were enriched in Tegument 1, e.g. genes expressed in > 90% of Tegument 1 cells and < 50% of Tegument 2 cells, including a Plexin domain-containing protein (Smp_348500 -Supplementary Fig. S4B). Remarkably, 12 genes out of the top 15 upregulated genes in Tegument 1 are uncharacterised proteins (Supplementary Table S6).
The unexpected overrepresentation of biological processes associated with 'nucleotide/side metabolism' in the sporocyst tegument prompted us to compare among the most significantly represented GO terms and top marker genes in the tegument of other developmental stages for which single-cell transcriptomic data are available15,16,17. The comparative analysis among the miracidium17, schistosomulum16, adult15 and sporocyst showed not only an expected GO term present across all stages (i.e., 'microtubule-based process'), but also biological processes found only in the sporocyst tegument, including purine 'ribonucleoside salvage', 'purine ribonucleotide salvage', 'nucleotide salvage', biological processes linked to nucleic acid metabolism as indicated above (Supplementary Fig. S5 and Table S9). The comparison between top marker genes in the tegument cluster(s) across different developmental stages, revealed genes that may be specific to the sporocyst tegument with tentative functions associated with the absorption of molecules, e.g., Smp_329690 described as nose resistant to fluoxetine protein 6; Smp_169090, a major facilitator superfamily (MFS) domain-containing protein (Supplementary Fig. S6 and Table S10).
Following the comparative analysis across developmental stages performed for the tegument clusters and to further understand the similarities between life stages and the consistency of marker genes, we extended this analysis to other tissues. This revealed conserved and tentative developmental stage-specific GO terms and top marker genes for stem cells (Supplementary Figs. S7, S8 and Tables S9, S11), parenchyma (Supplementary Figs. S9, S10 and Tables S9, S12), muscle cells (Supplementary Figs. S11, S12 and Tables S9, S13) and neurons (Supplementary Figs. S13, S14 and Tables S9, S14). It is important to highlight the limitations of this approach. The data obtained from each developmental stage were generated by different research teams, different versions of the genomes were employed for mapping, and different approaches were followed for the analyses (see Methods). Notwithstanding these caveats, several GO terms specific to the sporocyst tissues were identified, providing indications of tentative functional differences in this intra-molluscan stage compared to other developmental stages. For instance, the GO term 'cellular protein complex disassembly' is only found in neurons of sporocysts, but not in other developmental stages (Supplementary Table S9 and Fig. S13). Similarly, genes expressed in the sporocyst Parenchyma cluster were involved in carbohydrate, amino acid, lipid and iron metabolism, i.e., genes with tentative roles in the catabolism of nutrient molecules derived from the snail tissues. The enzymes UDP- glucose 4—epimerase (Smp_070780) and Ornithine aminotransferase (Smp_000660), the 14 kDa fatty acid-binding protein (Smp_095360), and Ferritin (Smp_063530, Smp_311630 and Smp_311640) were all within the top 30 marker gene list in the Parenchyma cluster of sporocysts, but not in the Parenchyma cluster(s) of the other developmental stages (Supplementary Figs. S9, S10 and Tables S9, S12).
Overall, these findings suggest that cluster-specific gene products display distinct functions in different molecular pathways, and that the identified cell populations may be involved in different biological processes in the mother sporocyst (Fig. 1C and Supplementary Tables S4 and S5). To spatially validate the predicted cell clusters, we defined highly specific cluster-defining marker genes (Fig. 2A and Supplementary Table S15), for which Fluorescence in situ Hybridization (FISH) probes were generated (Supplementary Table S2). We identified cells expressing the Muscle-specific marker myosin heavy chain, location of which correlated with actin filaments following the anterior–posterior axis of the sporocyst (Fig. 2B and Supplementary Fig. S15A, B). Whilst a Neuron-specific marker was expressed across a handful scattered cells in the mid region of the parasite (Fig. 2B and Supplementary Figu. S15C, D), cells expressing the Stem/germinal cluster-specific marker histone H2A were mainly located in clusters towards one pole (Fig. 2C). As stem cells are located in the posterior half of the miracidium larva17, we hypothesise this is the same in the mother sporocyst, so the Stem/germinal cell markers are thus likely to be highlighting the posterior end. In addition, a few individual Stem/germinal cells were located in the medial region towards the surface of the animal (Supplementary Fig. S15E, F). The two Tegument cell clusters that highly expressed the micro-exon gene 6 (MEG-6), were spatially validated by a strong FISH signal from cells lining the surface of the parasites (Fig. 2D, Supplementary Fig. S16 and Supplementary Video S1). The Parenchyma cluster cells were identified by the cluster specific expression of Smp_318890 (encoding a hypothetical protein). Even though this marker was expressed in < 50% of the cells (Fig. 2A), it showed a high and specific expression in parenchyma cells. These cells seem to be distributed throughout the whole parasite body with a tendency towards the posterior pole (Fig. 2E, Supplementary Fig. S15G, H, and Supplementary Video S2). The parenchymal cells showed clear anterior–posterior cytoplasmic projections containing Smp_318890 transcripts (Fig. 2E, yellow arrowheads).
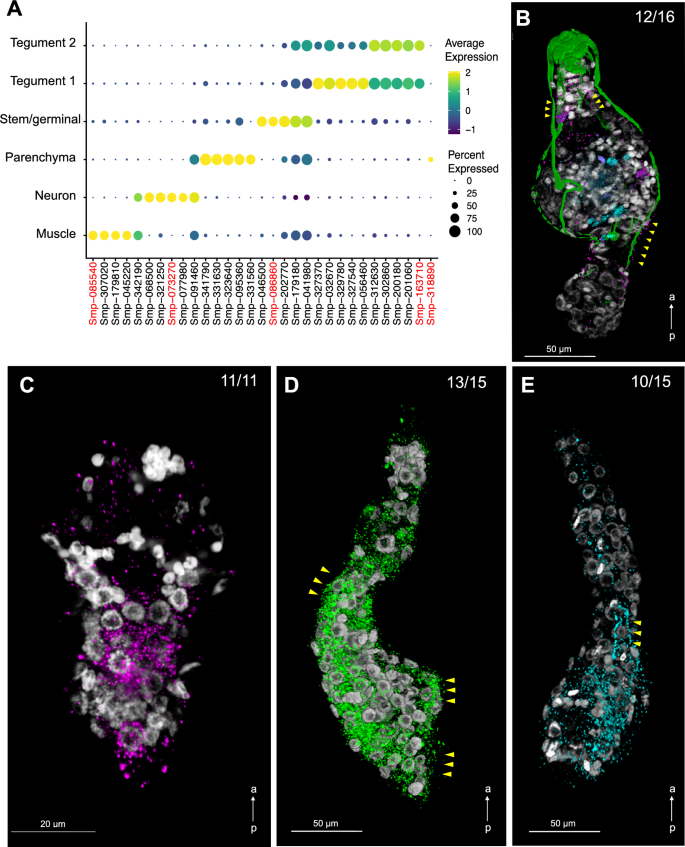
Cell clusters spatially validated by fluorescence in situ hybridization (FISH). (A) Dot plot showing the expression of the top 5 markers identified for each cell cluster. The average gene expression level for each marker is represented by a colour gradient from dark blue (low expression) to bright yellow (high expression). The circle sizes indicate the percentage of cells in each indicated cluster. The top marker genes for each cluster were defined as the highest AUC scores, which are calculated using FindAllMarkers (Seurat) using both presence and absence, and the level of expression. FISH probes for the following cluster-specific markers (highlighted in red) were used for spatial validation: Pan-tegument, micro-exon gene 6 or MEG-6 (Smp_163710); Muscle, myosin heavy chain (Smp_085540); Stem/Germinal, histone H2A (Smp_086860); Parenchyma, hypothetical protein (Smp_318890); and Neuron, neuroendocrine protein 7b2 (Smp_073270). Full data for the indicated top markers are provided in Supplementary Table S15. (B) Double FISH with myosin heavy chain (Smp_085540; magenta) and neuroendocrine protein 7b2 (Smp_073270; cyan) probes identified muscle and neuron cell clusters, respectively. Phalloidin-stained actin filaments are shown in green and DAPI staining in grey. Yellow arrowheads indicate co-localisation of myosin heavy chain and actin filaments. n = 16 parasites. (C) Localisation of the stem/germinal cells using FISH with histone H2A (Smp_086860; magenta). DAPI staining in grey. n = 11 parasites. (D) Localisation of tegumental cells using FISH with MEG-6 (Smp_163710; green). DAPI staining in grey. Yellow arrowheads indicate cells lining the surface of the parasites. n = 15 parasites. (E) Localisation of parenchyma cells using FISH with hypothetical protein (Smp_318890; cyan). DAPI staining in grey. Yellow arrowheads point to cytoplasmic projections containing Smp_318890 transcripts. n = 15 parasites. Scale bars: 50 μm in panels (B), (D), (E), and 20 μm in panel (C). a ← p: anterior–posterior axis.
Stem cell heterogeneity revealed by self-assembling manifold algorithm
Using the self-assembling manifold (SAM) algorithm24, the 119 stem/germinal cells were further analysed. Three discrete subclusters (clusters 0, 1 and 2) with distinct transcriptional profiles were identified (Figs. 3A, 3B and Supplementary Tables S16, S17). We used Scanpy25 to rank genes that characterise each of the stem cell subclusters (Supplementary Table S18) and the five top-ranked genes for each cluster were used for visualisation. STRING26 was used to predict protein–protein interactions among the top 50 ranked genes within each subcluster (Supplementary Table S19). The analysis of subcluster 0 showed a network with strong connectivity of inferred protein–protein interactions among the top marker genes (Figs. 3C). The top STRING terms for subcluster 0, across all categories, are related to ribosomes and translation (Supplementary Table S19). In contrast, weak connectivity was found between genes expressed in subclusters 1 and 2, with no network, far fewer STRING terms, and weaker statistical support (Supplementary Table S19).
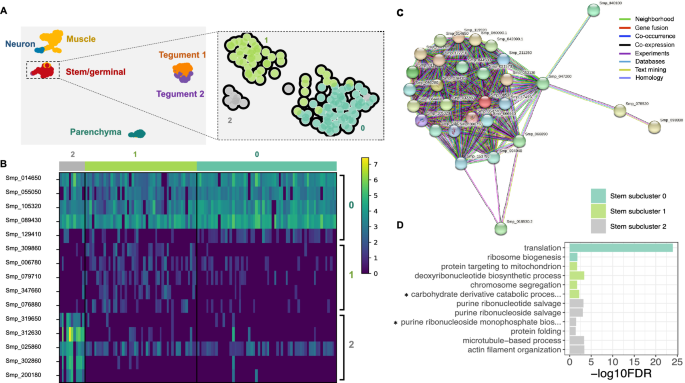
Stem/germinal cell sub-clusters. (A) Clustering of the sporocyst data using the self-assembling manifold (SAM) algorithm for all cells (left) and for the Stem/germinal cluster only (right). The SAM algorithm with Leiden clustering identified three stem/germinal subclusters (0, 1 and 2). The lists of SAM topology genes for all cell clusters or stem/germinal cell cluster only are provided in Supplementary Tables S16 and S17, respectively. (B) Heatmap of expression of the top 5 marker genes identified in each of the three stem/germinal subclusters. The average gene expression level for each marker is represented by a colour gradient from dark blue (low expression) to bright yellow (high expression). The full list of top marker genes identified by Scanpy in the three SAM stem cell subclusters is provided in Supplementary Table S18. (C) Interaction network analysis by STRINGdb for stem/germinal subcluster 0. The coloured nodes of the network represent proteins (genes ID for each protein are indicated, and all splice isoforms or post-translational modifications for each protein are collapsed). The coloured edges indicate sources of the interaction evidence as described. The full list of all enriched String terms for the top 50 markers of each stem/germinal sub-cluster is provided in Supplementary Table S19. (D) Stem/germinal subcluster gene ontology (GO) enrichment analysis in the category biological process, for top marker genes in each indicated colour-coded cell cluster. Only statistically significant GO enriched biological processes are depicted (− log10 (FDR < 0.05)), and only terms supported by ≥ 2 genes are shown. FDR: False discovery rate. Full names of GO terms indicated with *: carbohydrate derivative catabolic process (GO:1901136), purine ribonucleoside monophosphate biosynthetic process (GO:0009168). Full data provided in Supplementary Table S20.
Using TopGO we further explored the biological roles of cells within each stem/germinal subcluster by examining over-represented Gene Ontology terms. Related annotations from multiple genes within each cluster were evident; for example, and consistent with the STRING findings, subcluster 0 was highly enriched for biological processes 'translation' and 'ribosome biogenesis' (Figs. 3D and Supplementary Table S20). Statistically significant biological processes, supported by a minimum of two genes, were identified for the other two subclusters. In subcluster 1, 'deoxyribonucleotide biosynthesis' and 'chromosome segregation' suggest active DNA synthesis and cell division (Supplementary Table S20). In subcluster 2, 'actin filament organisation' and 'microtubule-based process', supported by three and six genes, respectively, as well as 'purine ribonucleoside salvage', 'purine ribonucleotide salvage' and 'purine ribonucleoside monophosphate biosynthetic process' indicate considerable similarity to the tegumental gene clusters described above (Supplementary Table S20).
Previously, three stem cell populations in mother sporocysts18 were identified as kappa, delta and phi based on the differential expression of seven marker genes (klf, nanos-2, fgfrA, fgfrB, p53, zfp-1, and hesl). Although these markers were expressed in our dataset, we were not able to use them to unambiguously assign the Stem/germinal cell subclusters identified by SAM to the kappa, delta and phi cell populations (Supplementary Figs. S17–S19).
Promoter motifs and transcription factor binding sites enriched in two major cell populations
To further investigate gene expression regulation in the Stem/germinal cell population, we searched for putative Transcription Factor Binding Sites (TFBSs). The Tegument-1 cell cluster was included as a control because it represents a somatic/differentiated cell population with a similar number of cells as the Stem/germinal cell population. Motif analysis was performed for marker genes specific to the Stem/germinal cluster (12 genes; Fig. 4A) and the Tegument-1 cluster (49 genes; Supplementary Fig. S20A). We identified five enriched motifs in the promoter region (i.e., 1 kb upstream the Transcription Start Site—TSS) of Stem/germinal cell marker genes, and ten in those of tegumental cells (Fig. 4B, Supplementary Fig. S20B and Table S21). Interestingly, nearly all enriched motifs had ≥ 1 significant match(s) to known TFBSs in the model worm C. elegans based on JASPAR 2022 database, and no overlapping motifs were identified between the two analysed cell clusters (Supplementary Table S22). Within both the Stem/germinal (Fig. 4C) and Tegument-1 cell clusters (Supplementary Fig. S20C), most marker genes have binding sites for multiple transcription factors (TFs). Notably, from the 12 Stem/germinal cell marker genes, 11 contain the motif S-STREME-1, that is similar to the binding site for ceh-22, a homeobox gene and orthologue of the human NKX2-2 gene (Fig. 4D). A further eight genes share the binding motif for pha-4 (S-STREME-5), a forkhead/winged helix factor, and eight genes share a motif (S-STREME-4) similar to the TFBSs for both ceh-22 and vab-7 homeobox genes (Fig. 4C). Six genes share S-STREME-2, which is similar to the unc-30 homeobox binding motif, and three genes share S-STREME-3, similar to the binding motif for the transcription factor sma-4 (SMAD/NF-1 DNA-binding domain factors; Supplementary Table S22). Detailed information of motif locations within the promoter region of Stem/germinal and Tegument 1 marker genes is provided in Supplementary Table S23.
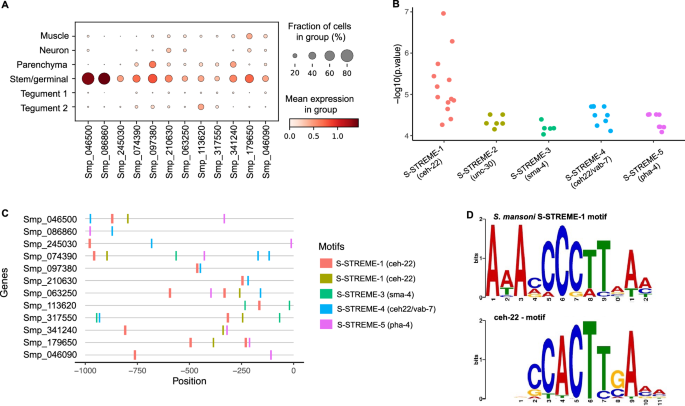
Promoter motif and transcription factor binding sites in stem/germinal cells. (A) Dot plot showing the expression level of the 12 Stem/germinal cell cluster-specific marker genes used for the analysis. Fraction of cells (%) and mean expression are indicated. The average gene expression level for each marker is represented by a colour gradient from white (low expression) to dark red (high expression). (B) Distribution of the − log10 (p values) for the top 5 ranked motifs identified in the 12 stem/germinal cell marker genes. The x-axis indicates motif names from XSTREME and significant match (p < 0.05) to known Transcription Factors Binding Sites (TFBSs) in the JASPAR 2022 nematode dataset (https://jaspar.genereg.net/downloads/). The y-axis represents log-transformed p values of each motif site shown in C. Full data provided in Supplementary Tables S21 and S22. (C) Predicted position distribution of the top 5 ranked motifs along the promoter region of the stem/germinal cell marker genes. The promoter region was taken as 1 kb upstream of the Transcription Start Site (TSS). Full data provided in Supplementary Table S23. (D) TF binding motifs found enriched in the promoter region of stem/germinal cell cluster marker genes. Schistosoma mansoni enriched motif named S-STREME-1 with the sequence: 1-AAAMCCCTTAAM (top) found in 11 of the 12 stem/germinal cell cluster marker genes with significant match to the binding site MA0264.1 (MA0264.1.ceh-22) for C. elegans ceh-22 (bottom) in the JASPAR database (https://jaspar.genereg.net/). The height of the letter in the motif scheme represents the frequency of the nucleotide observed in each indicated position.
Using the above TFs from C. elegans, we identified putative S. mansoni orthologs. The putative S. mansoni TFs were examined by KEGG ortholog search, and the binding profiles were further confirmed for some candidates using the Jaspar Profile Inference tool27. This screening produced a list of 12 candidate S. mansoni TF genes (Supplementary Table S24): one ortholog of sma-4 (Smp_033950); two orthologs of ceh-22 (Smp_027990 and Smp_186930), pha-4 (FoxA) (Smp_331700 and Smp_158750) and unc-30 (Smp_124010 and Smp_163140); and five orthologs of vab-7 (Smp_147640, Smp_138140, Smp_347890, Smp_308310, Smp_134690). Two of the 12 candidates (Smp_186930 and Smp_033950) were not annotated as TFs in the KEGG search but indicated as orthologues to C. elegans counterparts. Based on a meta-analysis of gene expression for S. mansoni developmental stages28, the Stem/germinal cell cluster specific markers (Fig. 4A) with promoter motifs and TFBSs identified above, tend to show a high expression in the ovary of mature females, miracidia and sporocysts (Supplementary Fig. S22A). Interestingly, the sma-4 ortholog Smp_033950 (Mothers against decapentaplegic or MAD homolog) also showed a high expression across these developmental stages, particularly in sporocysts, relative to the other stages (Supplementary Figure S22A). At the single-cell level, this gene was expressed across all cell clusters, with the highest expression and percentage of cells in the Neuron cluster (Supplementary Figure S22B). Even though the TF Smp_033950 was expressed in < 5% of the stem/germinal cells (Supplementary Fig. S22B), it was the only one among the 12 putative TFs identified in this study (Supplementary Table S22) that showed a Stem/germinal cluster-specific expression and almost exclusively in subclusters 1 and 2 (Supplementary Fig. S22C).
To explore the evolutionary conservation of the predicted of TFBSs within the promoter region of Stem/germinal and Tegument cell cluster marker genes, we searched for enriched common regulatory binding sites across seven Schistosoma species in addition to S. mansoni (Fig. 5A and Supplementary Fig. S21A). Several conserved motifs and combinations of motifs were significantly enriched within 1 kb upstream of the TSS of S. mansoni marker gene orthologues across all the analysed Schistosoma species (Fig. 5B and C, Supplementary Fig. S21B and Supplementary Tables S25–S27). Moreover, the closer the species in the phylogenetic tree, the more conserved the sets of promoter motifs for each marker gene.
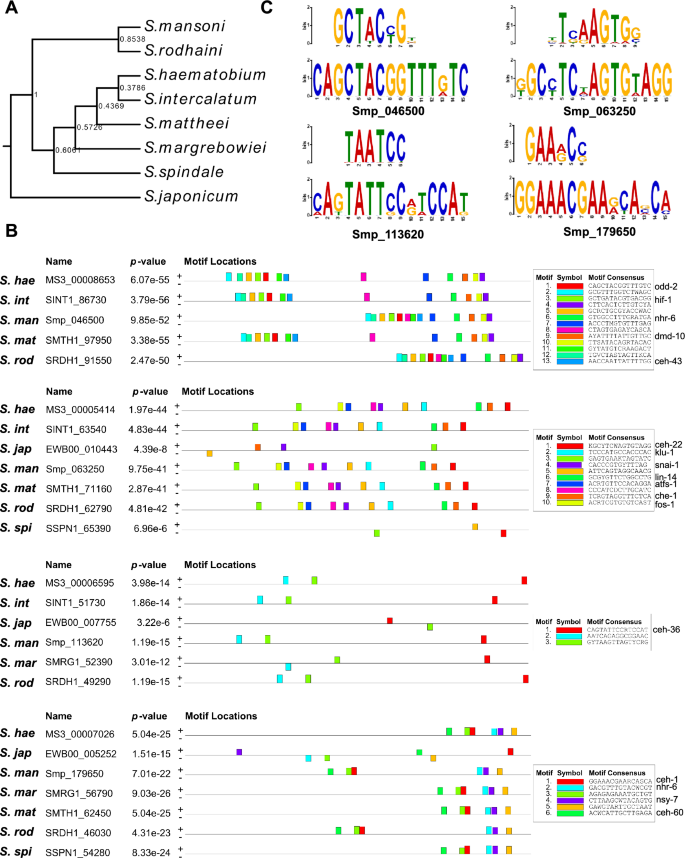
Promoter motif conservation of stem/germinal cell genes in the Schistosomatidae family. (A) Rooted species tree inferred with Orthofinder, showing the phylogenetic relationship between the indicated species. Branch support values are indicated. (B) Motifs found enriched in the promoter region of the orthologs in Schistosomidae to the S. mansoni marker genes Smp_046500, Smp_063250, Smp_113620 and Smp_179650. The full list of analysed orthologous genes is provided in Supplementary Table S25. The colour-coded motifs detected for each group of orthologs are indicated along the 1 kb region upstream of the Transcription Start Site (TSS) for each gene. Significant matches with binding site found for C. elegans in the JASPAR database (https://jaspar.genereg.net/) were annotated. Full data provided in Supplementary Table S27. S. hae: S. haematobium, S. int: S. intercalatum, S. man: S. mansoni, S. mat: S. mattheei, S mar: S. margrebowiei, S. rod: S. rodhaini, S. spi: S. Spindale, S. jap: S. japonicum. (C) Logo of most significant motifs identified for each group of promoter regions of orthologous genes and the corresponding binding site for C. elegans in the JASPAR database (https://jaspar.genereg.net/). Sequence CAGCTACGGTTTGTC (bottom) found in orthologs of Smp_046500 showed a significant match to the binding site MA2148.1 (MA2148.1.odd-2) for C. elegans odd-2 (top); sequence KGCYTCWAGTGTAGG (bottom) found in orthologs of Smp_063250 showed a significant match to the binding site MA0264.2 (MA0264.2.ceh-22) for C. elegans ceh-22 (top); sequence CAGTATTCCRTCCAT (bottom) found in orthologs of Smp_113620 showed a significant match to the binding site MA2159.1 (MA2159.1.ceh-36) for C. elegans ceh-36 (top); sequence GGAAACGAARCASCA (bottom) identified in orthologs of Smp_179650 showed a significant match to the binding site MA0260.1 (MA0260.1.che-1) for C. elegans che-1 (top). The height of the letter in the motif scheme represents the frequency of the nucleotide observed in each indicated position.
In addition to the predicted TFs and TFBSs expressed in Stem/germinal cluster and Tegument clusters, we investigated in our dataset the expression of well-characterised TFs in other developmental stages29. The expression of the flatworm-specific zinc finger proteins genes zfp-1 and zfp-1-1, involved in the tegument specification in adult worms, was detected in few sporocyst cells (< 20) in the Stem/Germinal cluster (in the three subclusters—Supplementary Table S18) and in the Tegument (in particular in Tegument 2—Supplementary Table S3), respectively (Supplementary Figure S23).
Discussion
In this study, we employed single cell transcriptomics, followed by spatial validation, to unveil the cellular components of the mother sporocyst, the life cycle stage that marks the start of rapid asexual proliferation within the intermediate host. Due to the experimental challenges of studying parasites within snail tissues, we have used mother sporocysts cultured in vitro for 5 days, an approach that has been shown to mirror many aspects of in-vivo parasite development in snails30,31,32,33. For instance, the first scRNA-seq study in schistosomes described the transcriptomic signatures of proliferating cells derived from mother sporocysts transformed in vitro18. The authors validated their findings in parasites developing in vivo, following parasite progeny through genesis of cercariae and intra-mammalian stages18. Molecular and cellular differences cannot be completely ruled out; however, in-vivo validation of observations across several stages of development18 provides strong evidence that the in-vitro miracidium–sporocyst transformation, and culture of early sporocyst stages, is a reliable model to study the parasite development.
Striking changes have been reported during the miracidium–sporocyst transition, including shedding of the ciliary plates, tegumental remodelling and an overall tissue reorganisation21 that transforms the free-living miracidium with ~ 365 cells17 into a simple 'sac-like structure' enriched with stem/germinal cells10. With the assistance of a novel machine learning approach22, we estimated an average of 169 nuclei in a single D5 mother sporocyst (ranging from 112 to 254 nuclei) and have revealed previously uncharacterised variability in cell number among the parasites at ...
Comments
Post a Comment